Biophysics: Biopolymer Networks & Brownian Dynamics
Modeling of the cytosceleton
Kei Müller, Christoph Meier, Dhrubajyoti Mukherjee and Christian Cyron
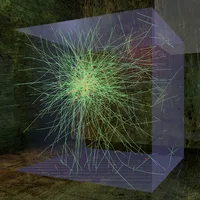

Mechanical stability as well as many metabolic activities of eucariotic cells are to a large extent owed to a fibrous network of proteins (e.g. Actin and Myosin) called the cytoskeleton.
It plays a pivotal role in processes such as cell migration, shape change or cell division. Though being of great importance for the overall functionality of a cell, many properties of the cytoskeleton remain unknown or only partly understood.
By the means of numerical simulation, we study the formation of such biopolymer networks and their varying phenotypes. Furthermore, we are interested in the viscoelastic properties of these networks as a deeper understanding of the cytoskeleton's mechanics would eventually help to shed light on the origins of its enormous variability and flexibility. This knowledge may support or even drive the development of new methods in fields such as tissue engineering or cancer therapy.
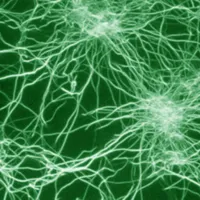
The pictures on the right show cluster networks which were observed in vitro and found via simulation. Simulations are conducted using finite beam elements. An initially entangled solution of filaments interacts with a second molecule species, the crosslinkers, and thus forms the displayed network. Filament and crosslinker motion is driven by thermally induced random external forces.
The following movie shows the formation of a layer-type actin network which results from adding a specific kind of crosslinking molecules to the simulated volume.
Publications
Please find publications on this topic here.